iTRAQ / TMT Labeling Quantitative Proteomics
Introduction:
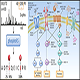
iTRAQ (4 plex or 8 plex) and TMT (2 plex through 10 plex) are chemical labeling methods that enable high through-put and internally controlled quantifications in proteomics studies. This is another very popular service in Omics Technologies which enables researchers to directly compare the difference of proteomes in their treated vs untreated, diseased vs normal, over-expressed vs control, gene silenced vs control, knockout vs wild type, mutant vs wild type groups, etc.
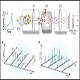
Every sample group in the study is normalized to start from the equal amount of total protein/peptide. Protein digest of each sample is chemically labeled with a sample specific tag. Tagged peptides from different samples are pooled together with equal total individual amount and the complete catalogue of proteins in the pooled sample is resolved by Mass Spectrometry analysis and each protein identified in the study will contain 4 or 8 (for iTRAQ) and 2 through 10 (for TMT) intensity values representing this protein's abundance in different samples. The result shows not only the identification of peptides/proteins, but also the absolute/relative abundance of the peptides/proteins in each sample.
Your samples (several test groups and control group) will be digested with Trypsin and peptides are chemically labeled by either iTRAQ or TMT labeling method, and the labeled peptides are pooled and ion-exchanged through Strong Cation Exchange (SCX) HPLC cleaning. To improve the sensitivity, we suggest the basic Reverse Phase LC (bRPLC) to further fractionate your cleaned sample into 96 fractions and then pooled to 24 fractions to maximize the coverage and depth of your analysis. 24 Mass Spectrometry Analysis runs will be performed by the world's most advanced Orbitrap LC-MS/MS system.
There are typically 10 steps in a chemical labeling Omics Technologies quantitative proteomics project:
1) Sample preparation – Cell or tissue lysis, biofluid (plasma or serum) depletion; protein quantification on all samples.
2) Sample digestion – Reduction and alkylation followed by in-solution trypsin digestion and C18 cleaning before labeling.
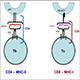
3) Sample labeling – Isotope labeled reagents will be applied to label 2 through 10 samples individually simultaneously.
4) Sample pooling – Equal amounts (according to total protein quantification) of labeled samples are pooled together to allow unbiased downstream purification and Mass Spectrometry Analysis.
5) SCX cleaning – Ion-exchange to remove excessive chemicals and fractionation of pooled sample through off-line SCX HPLC.
6) 2D fractionation – bRPLC separation of the SCX cleaned samples into 96 fractions, and further pooled into 24 fractions based on the rule of avoiding co-elution.
7) Nano LC/MS/MS – 50 nL/min liquid chromatography to enable super high ionization efficiency for downstream Mass Spectrometry Analysis using the state-of-the-art Orbitrap Mass Spectrometer. 24 fractions will be analyzed in 24 Mass Spectrometry runs.
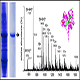
8) Protein profiling – Protein database searching with Mascot.
9) Data analysis – Data validation, visualization and quantification using commercial softwares and Omics Technologies's unique R packages and scripts.
10) Report generation – Report will be sent to you in Excel format as well as a summary in PDF format. We will also provide you any details you need for your papers' MATERIALS AND METHODS section. We will make sure you understand your result and help you with your paper writing with free follow-up services.
Sample types we accept:
1, 2D gel spots, SDS-PAGE bands
2, Cell Lysates and Tissue Lysates
3, Biofluids, such as plasma*, serum*, saliva, tear, etc.
*We provide High Abundance Protein Depletion Service to significantly (100-500 folds) increase the depth of your biofluid proteomics analysis by removing top abundant proteins from your samples. Read more for details.
4, FFPE slide/ FFPE extract
5, Customized sample types (please contact us to discuss)
Please refer to our Omics Technologies Proteomics Sample Preparation and Shipping Guidelines for instructions on how to prepare your samples with less contamination for Mass Spectrometry Analysis.
-
Phone:
+1 888-206-0066 (TOLL FREE) -
Email:
info@omicstech.com